Chemistry
Atomic Orbitals
In Quantum Mechanics, Atomic Orbitals are mathematical functions that describe the wave-like behavior of electrons in atoms. This function can be used to calculate the probability of electrons appearing around the nucleus, and the meaning of “orbital” refers to the probability of electrons appearing in a specific area. According to the “shape” of the track, it can be classified into s, p, d, f, etc.
Electronegativity
Electronegativity describes the ability of atoms of an element to attract electrons in a compound. The greater the electronegativity of an element, the stronger the ability of its atoms to attract electrons in the compound. In a period of the periodic table, the electronegativity of the element atom increases from left to right; and it decreases from top to bottom in a group. Therefore, the elements at the upper right of the periodic table (O, N, F, Cl, etc.) have higher electronegativity values. The element with the greatest electronegativity is fluorine.
Chemical Bond
A chemical bond refers to the strong interaction between atoms, ions, and other particles. Through chemical bonds, particles can form polyatomic compounds (such as organic molecules, inorganic molecules, ionic compounds, etc.). Simply put, for a polyatomic system, the most stable configuration between positively charged nuclei and negatively charged electrons is that when electrons are located between nuclei, electrons are attracted between different nuclei, and using this force the nuclei are “attracted” together, forming a chemical bond.
Ionic Bond: A chemical bond formed by electrostatic interaction between oppositely charged anions and cations, without directionality, such as sodium chloride (salt), calcium carbonate.
Covalent Bond: A chemical bond formed by sharing electron pairs between atoms. Two atoms with similar electronegativity are equally attracted to electrons, so they mainly form chemical bonds by sharing each other’s outer valence electrons. Covalent bonds are directional, resulting in complex molecular structures. For example, in the methane molecule, carbon atoms and hydrogen atoms are connected by covalent bonds to form a regular tetrahedron, the carbon atom is located at the center of the tetrahedron, and the hydrogen atom is located at the vertex of the tetrahedron. According to the number of shared electron pairs, it can also be classified into a single bond, double bond, and triple bond.
Hydrogen Bond: When a hydrogen atom forms a covalent bond with an atom with high electronegativity X (usually O, N, F), if it bonds with another atom with high electronegativity. When Y (usually also O, N, F) is close, using hydrogen as the medium between X and Y, a special form of interaction like X-H· · ·Y is generated, known as a hydrogen bond. Hydrogen bonds widely exist in biological macromolecules such as water and proteins and DNA. It plays a crucial role in stabilizing the conformation of biological macromolecules.
Functional Group
Functional groups are atoms or groups of atoms that determine the properties of organic compounds. Common functional groups include hydroxyl (-OH), carboxyl (-COOH), ether bond (C-O-C), carbonyl (C=O), halogen atom (-F, -Cl, -Br, -I), etc.
Aromatic
Aromaticity is a chemical property that exists in cyclic planar molecules co ntaining \(\pi\) bonds composed of delocalized electrons, which can provide molecules with stability that cannot be explained by conjugation alone. The number of electrons in the delocalized \(\pi\) of an aromatic molecule needs to satisfy the Huckel rule (also called the “4n+2” rule). Molecules with aromaticity are called aromatic compounds, and molecules without aromaticity are called aliphatic compounds. Aromatic compounds can be roughly classified into simple aromatic compounds (such as benzene), polycyclic aromatic compounds (such as naphthalene, and anthracene), and heterocyclic compounds (such as pyridine, and pyrrole).
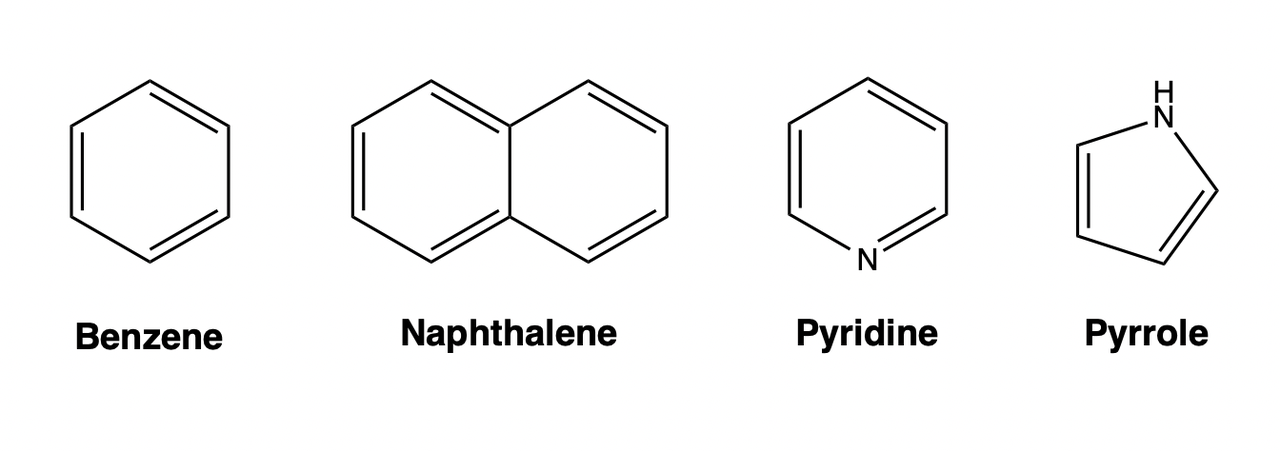
Conformation
Conformer usually refers to three-dimensional conformation, which refers to the structure that a molecule has in three-dimensional space. For organic molecules, their conformations cannot be randomly generated due to the limitation of the directionality of covalent bonds.
Isomers
In Organic Chemistry, substances with the same chemical composition (molecular formula) but different structures are called isomers of each other. For example, the compositions of ethanol and dimethyl ether are both \(\mathrm{C_2H_6O}\), but their structures are different:
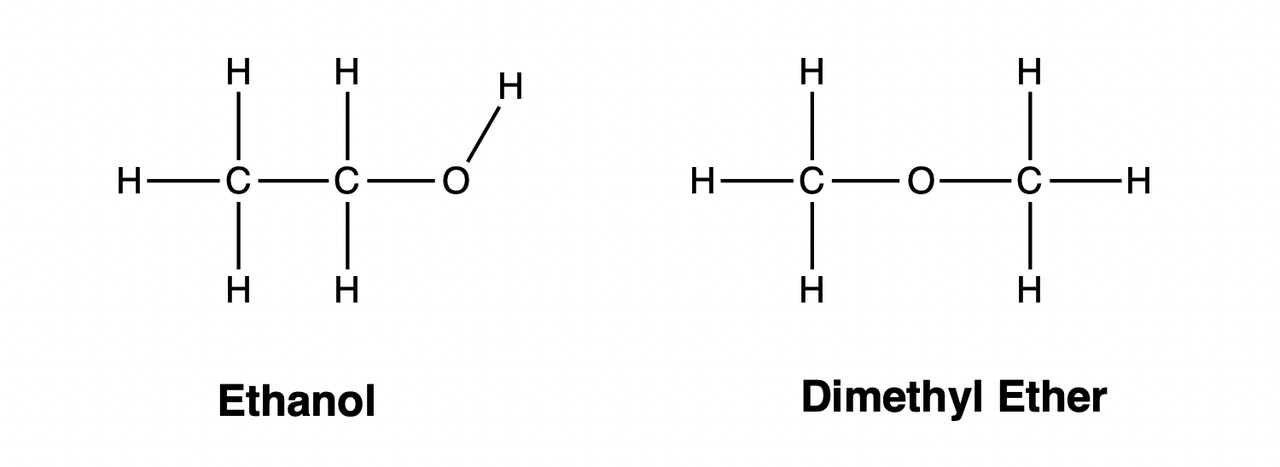
Stereoisomerism
Stereoisomers refer to molecules in which atoms are topologically connected in the same way but spatial arrangement of the atoms are different. For example, a molecule is likeley to have stereoisomers when it contains carbon atoms to which four different functional groups are bonded. Such atom is called chiral atoms, and usually R/S are denoted to distinguish two different them. In terms of biomolecules, such as peptides, amino acids and sugar, L/D are frequently used to denote different type of stereoisomers. The two amino acid configurations shown in the figure below are stereoisomers of each other. All natural amino acids are in the L configuration, and their carbon atoms are in the S configuration.
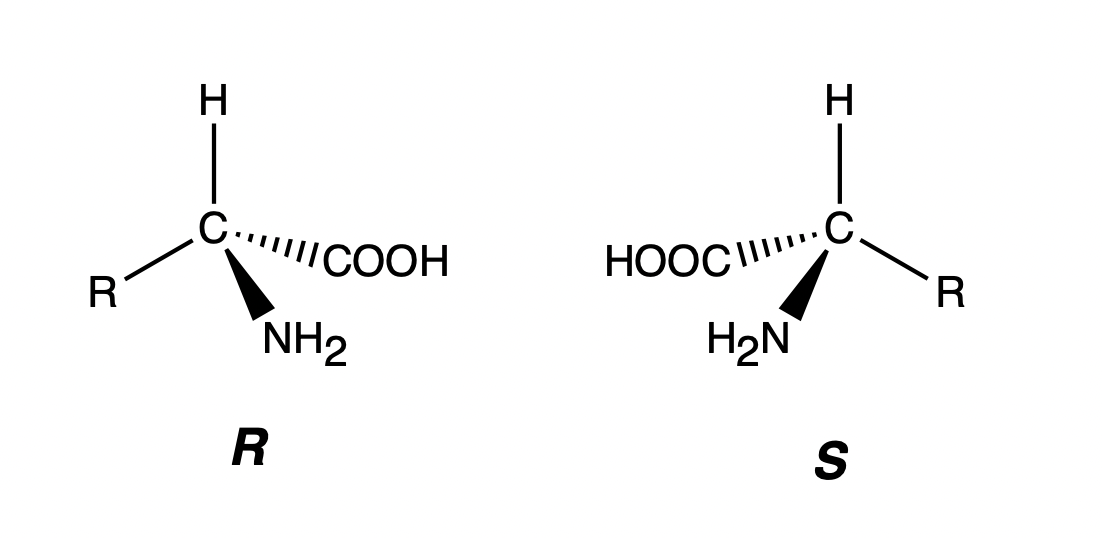
Cis-trans Isomerism
Cis-trans isomerism refers to isomerism that occurs due to the hindered free rotation in the compound molecule, which is commonly found in compounds with double bonds or rings.
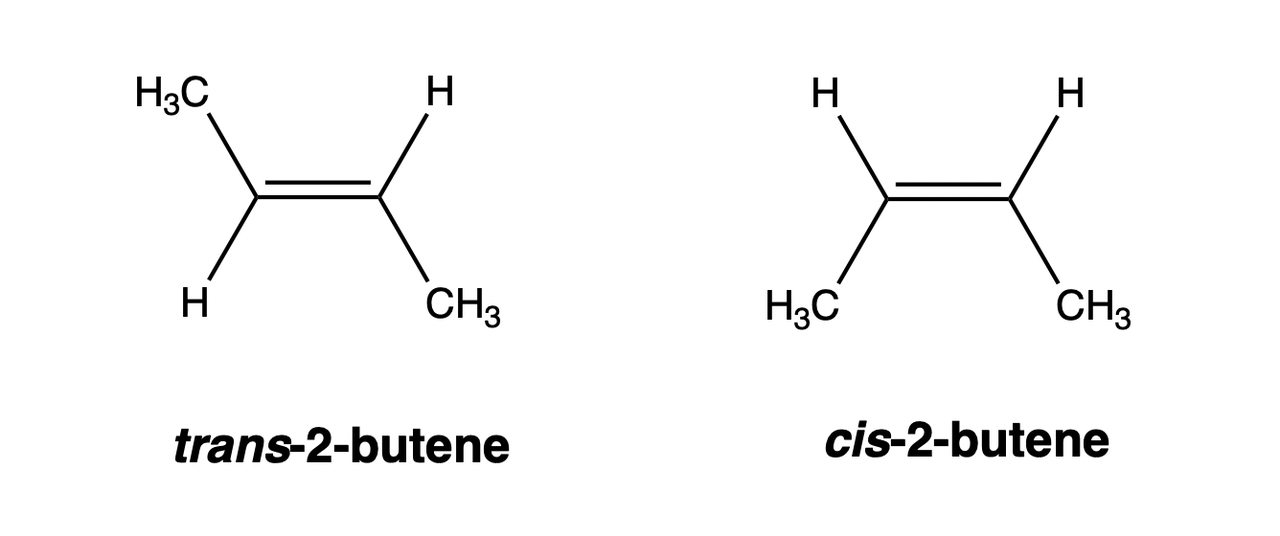
Tautomerism
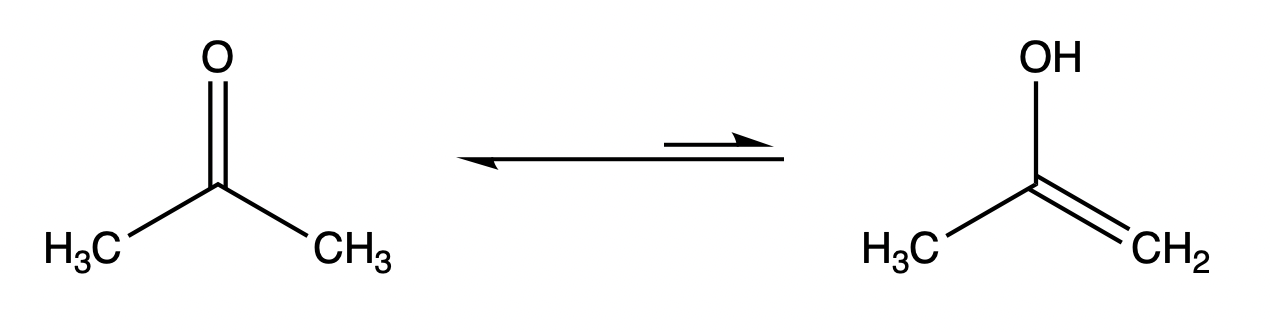
Tautomerism means the structure of some organic compounds is converted between two functional isomers. Most tautomerisms involve the transfer of hydrogen atoms or protons, and the conversion of single bonds to double bonds. The distribution of tautomers in equilibrium depends on specific factors, including temperature, solvent, and pH, etc. The diagram below shows the keto (left) and enol (right) tautomers present in carbonyl compounds, with the keto structure predominating in the usual case.
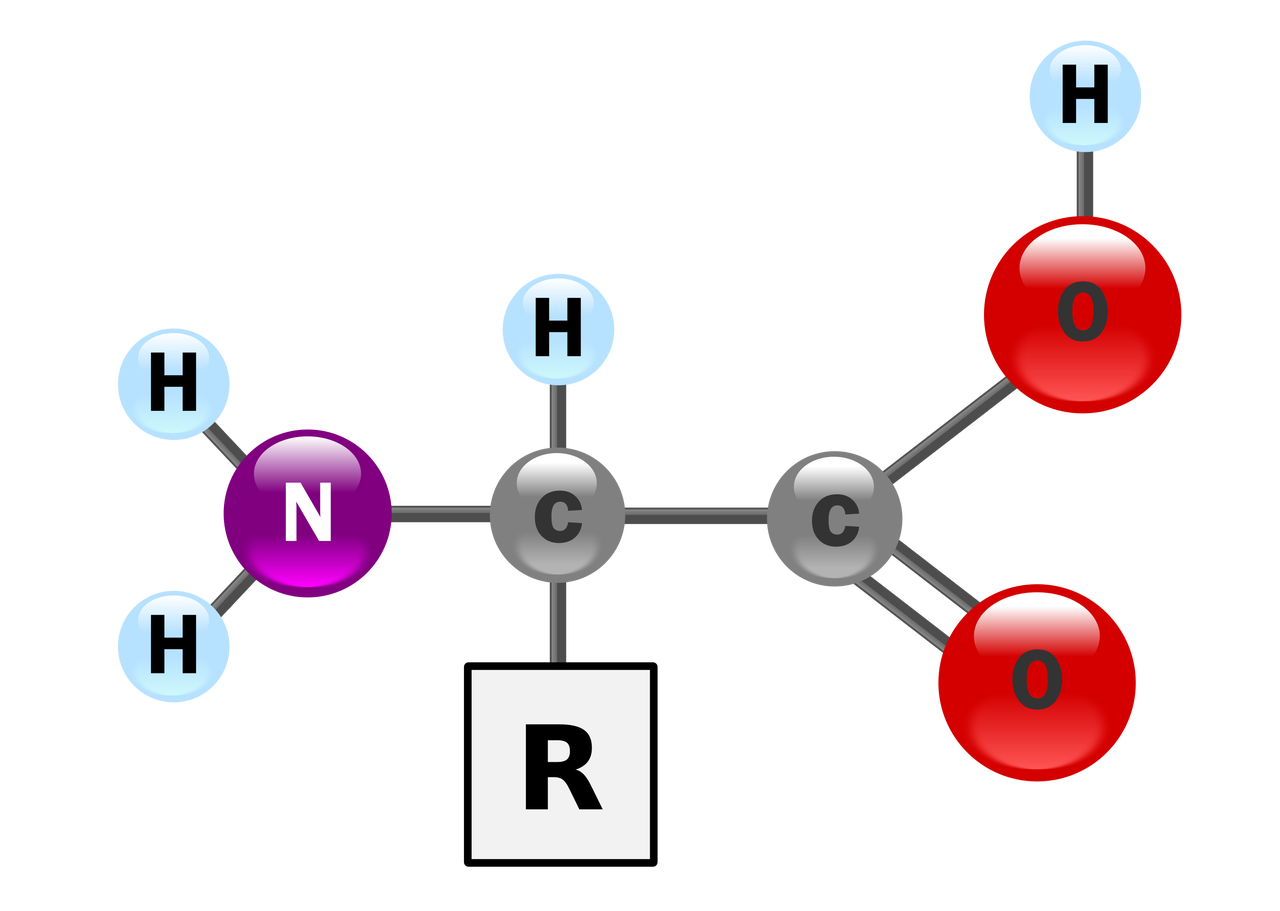
Amino Acids
Amino acids are biologically important organic compounds consisting of amino (-NH2) and carboxyl (-COOH) functional groups and side chains attached to each amino acid. Amino acids are the basic units that make up a protein. In nature, there are 20 genetically encoded amino acids.
Protein Structure
Protein structure refers to the spatial structure of a protein biomolecule, which can be divided into four levels to describe different aspects.
Primary structure: the linear amino acid sequence that makes up the polypeptide chain of a protein.
Secondary structure: a stable structure formed by hydrogen bonds between C=O and N-H groups between different amino acids, mainly \(\alpha\)-helix and β-sheet.
Tertiary structure: the three-dimensional structure of a protein molecule is formed by the arrangement of multiple secondary structural elements in three-dimensional space.
Quaternary structure: used to describe the interaction of different polypeptide chains (subunits) to form functional protein molecules.
Ligand
In biochemistry or pharmacology, a ligand refers to a compound that can bind to a receptor and then lead to some physiological effect. In medicinal chemistry, ligands are usually small organic molecules or short peptides composed of several amino acids. The forces between ligands and receptors are usually non-covalent interactions: such as hydrogen bonds, electrostatic interactions, van der Waals interactions, etc.
Receptor
Signal transduction is responsible for intracellular communication via series of molecular events (protein phosphorylation) upon chemical/physical signal outside cell , where receptor function in the central role as transmit signals outside cells and produce specific effects within cells. It is usually biological macromolecule such as protein. After the receptor binds to a specific stimuli, the structure will change to a certain extent, and the corresponding effect will be induced in the cell. In medicinal chemistry, receptors usually refer to target proteins able to bind with ligands.
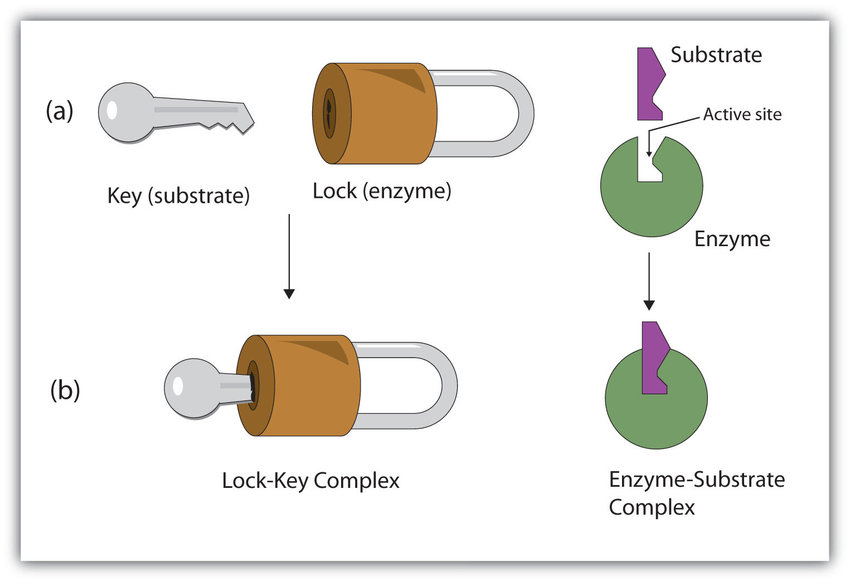
Lock and Key Model
The lock-and-key model is a theory proposed by E. Fischer in 1890 to explain the specific binding between enzymes and substrates (or between ligands and receptors). The model believes that the structures of enzymes and substrates at their binding sites should be strictly matched and highly complementary, just like the structural complementarity and matching of a lock and its original key. The disadvantage of this model is that the model treats the structure of the enzyme and the substrate as rigid structures, which is inconsistent with the fact that the conformation of the enzyme and the substrate changes during the catalytic reaction.
Induced Fit Model
The Induced-Fit Model is a model proposed by Koshland in 1958 to describe the enzyme-substrate (ligand-receptor) binding interaction. This model believes that in the process of binding the enzyme to the substrate, the substrate can induce a certain change in the structure of the enzyme, and finally form an active conformation that can bind to the substrate.
Molecular Docking
Molecular Docking is a technique that simulates the interaction between ligands and receptors. The technology predicts ligand binding modes and ligand-receptor binding forces by physically modeling intermolecular interactions and applying optimization algorithms such as the Monte Carlo method.
Reversible Reaction
A reversible reaction is a chemical reaction that can proceed in both the forward and reverse directions under the same conditions. When the degree of the reverse reaction direction is much smaller than that of the forward reaction direction, the reaction can be considered irreversible. Most of the reactions are reversible, such as the dissociation of weak acid/base, ligand-receptor binding, etc.
Chemical Equilibrium
Chemical Equilibrium refers to a state in which the forward and reverse reaction rates of a chemical reaction are equal in a reversible reaction with certain macroscopic conditions, and the concentrations of the reactants and the components of the products do not change. Take the following reaction as an example:
When the equilibrium is reached, the concentrations of \(\mathrm{A,B,C}\) are respectively [\(A\)],[\(B\)],[\(C\)], then the equilibrium constant K can be defined:
Given the reaction conditions, the equilibrium constant for a reaction with a fixed stoichiometric ratio is the same, and is related to the free energy change of the reaction as follows:
van der Waals force
van der Waals (vdW) force refers to the non-directional, unsaturated, weak interaction force between atoms. Van der Waals interactions are much weaker than chemical bonds, but they will significantly affect the melting point, boiling point, and many other properties. Van der Waals interactions have 3 major contributions:
Attractive or replusive interactions are between permanent charges, dipoles, quadrupoles, etc.
Induction (also known as polarization), which is the attractive interaction between a permanent multipole on one molecule with an induced multipole on another. This interaction is sometimes called Debye force.
Dispersion (usually named London dispersion interactions after Fritz London), which is the attractive interaction between any pair of molecules, including non-polar atoms, arising from the interactions of instantaneous multipoles.
In molecular simulations, van der Waals forces are usually described in terms of the Lanner-Jones potential function, which has the following form:
Where \(r\) is the distance between two atoms, \(C\) is a parameter, usually obtained by fitting physical quantities such as density and the enthalpy of evaporation.
Hydrophobic interaction
Hydrophobic interaction, also known as a hydrophobic effect, is a chemical phenomenon that which groups with hydrophobicity in an aqueous solution (such as alkyl groups without polarity) are close to each other to reduce the contact area with water. Hydrophobic interactions are the main driver of protein folding.
Thermodynamics
Thermodynamics focuses on the interaction of heat and work between chemical reactions and system states under the laws of thermodynamics. Generally speaking, the problems (equilibrium state) that do not involve the study of the chemical reaction process belong to the category of chemical thermodynamics, such as phase transition, and the balance of sodium and potassium ions on two sides of the cell membrane.
Kinetics
Kinetics, also known as reaction kinetics and chemical reaction kinetics, is a branch of physical chemistry that studies the rate and mechanism of chemical reactions. Chemical kinetics is different from chemical thermodynamics. It does not care about the equilibrium state, but studies the chemical reaction dynamically, and studies the time required for the transformation of the reaction system, as well as the microscopic process involved.